This might be replaced by an Interface in the future. This can be done by accessing the JmolViewer class. It is also possible to get data back out of Jmol. This is nicely documented by the Integration.java example. Most interaction with Jmol will happen by sending RasmolScript - like commands to Jmol. Another example: Programmatic Access to Jmol.A special page is devoted to using Jmol as 3D viewer for CDK based projects: Jmol Cdk Integration.To see how Spice is integrating Jmol, please have a look here.

#Jmol simulation code
Here is the previous code ported to Jython.
A very good start is the Integration.java out of Jmol SVN.TouchMol - molecular visualization on a multi-touch table, allowing up to 4 users to interact with a molecular model using their fingers.STRAP - Alignment Program for Proteins and workbench for protein structures.A java webstart version can be run from online. Spice - Spice is a DAS client for distributed annotation of protein sequences and structures.It uses Jmol for its 3D interactive plotting. Sage is a computational platform with the goal of providing a viable free and open-source alternative to Matlab, Maple, Mathematica and Magma.Raman Data Search and Storage (RDSS) A freeware and user-friendly application developed as an analytical tool for a fast and accurate identification of unknown minerals by comparison of their Raman spectra with the indexed library of data.
#Jmol simulation for mac os x
It is a dashboard application for Mac OS X that uses the JmolApplet. ProteinGlimpse is a free widget for visualizing macromolecules retrieved from the Protein Data Bank or from local disk.PFAAT - Protein Family Alignment Annotation Tool.PMOL - A Processing sketch using JMOL to load Molecules, and Processing to visualize them.Molecular Workbench - A Molecular Simulation Tool.MoCalc2012 is a Graphical User Interface for MOPAC, GAMESS(US), Firefly and ORCA that uses Jmol for displaying geometries, orbitals, surfaces, animations and vibrations using the Jmol scripting language.Janocchio is an applet and application in addition to using Jmol display capabilities, it calculates both H-H and H-C 3-bond NMR coupling constants and NOEs from a three-dimensional structure.J-ICE can deal with CASTEP, CRYSTAL09 (as well as 06, 03 and 98), QUANTUM ESPRESSO, VASP, Wien2k, FHI-aim, CIF, PDB and many others formats.
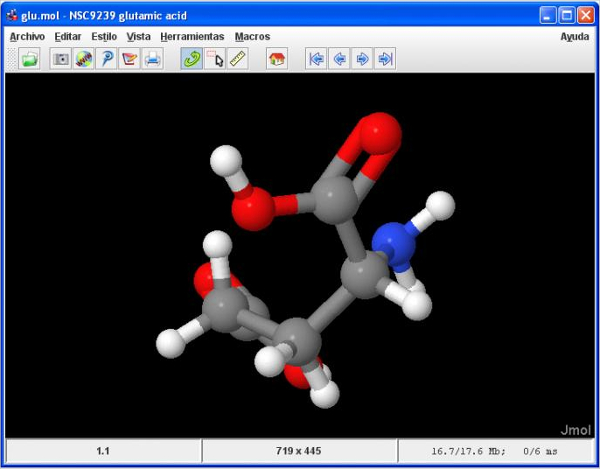
#Jmol simulation software
(There is another page for operating systems and software suites that include Jmol.) Jmol users, please add here your favorite Java applications that embed Jmol. Course Management Systems, Learning Management Systems, Virtual Learning Environments, e-Learning Platforms
